\documentclass[twocolumn]{article}
\usepackage{harvard}
\usepackage{bibtexlogo}
\usepackage{html}
\usepackage{epsfig}
\usepackage{bmcmtr}
\sloppy
\begin{latexonly}
% Restrict info issued for underfull boxes:
\showboxbreadth=0
\showboxdepth=0
\end{latexonly}
\newcommand{\hlnk}[2]{\htmladdnormallink{#1}{#2}}
\newcommand{\hoturl}[1]{\hlnk{\texttt{#1}}{#1}}
\newcommand{\mailto}[1]{\hlnk{\texttt{#1}}{mailto:#1}}
\begin{latexonly}
% Fix hyphenation(s):
\hyphenation{McMullin}
\end{latexonly}
\newcommand{\hole}{\texttt{H}}
\newcommand{\sbst}{\texttt{S}}
\newcommand{\cat}{\texttt{K}}
\newcommand{\lnk}{\texttt{L}}
\newcommand{\lnkplus}{\mbox{\lnk$^{+}$}}
\newcommand{\blnk}{\texttt{BL}}
\begin{document}
\begin{latexonly}
\copyrightdate{1997}
% This must be manually echoed in the .latex2html-init
% also - l2h cannot (perversely) pick it up from here
% in time to put it in the HTML footer...
\end{latexonly}
\newcommand{\dcunumber}{\texttt{bmcm9702}}
\newcommand{\sfinumber}{\texttt{97-01-002}}
\title{SCL: An Artificial Chemistry in \emph{Swarm}}
\author{\htmlref{Barry McMullin}{mcmullin}\\
\mailto{mcmullin@eeng.dcu.ie}}
\date{\htmlref{\copyright
\thecopyrightdate}{copyright}}
\begin{titlepage}
\vspace*{\fill}
\maketitle
\thispagestyle{empty}
\vspace{10em}
{\large
\begin{center}
\hlnk{\textbf{Santa Fe Institute}}{http://www.santafe.edu}\\
Working Paper Number: \sfinumber{}
\end{center}
\vspace{4em}
\begin{center}
\hlnk{\textbf{Dublin City University}}{http://www.dcu.ie}\\
\hlnk{School of Electronic Engineering}{http://www.eeng.dcu.ie}\\
Technical Report Number: \dcunumber{}
\end{center}}
\vspace*{\fill}
\end{titlepage}
\begin{abstract}
This report describes the SCL (v0.04) system. This
is an implementation of an artificial
chemistry, using the
Swarm\footnote{\hoturl{http://www.santafe.edu/projects/swarm}}
simulation system. This chemistry is qualitatively
based on the system first described in
\citeasnoun{Varela:Autopoiesis}. This involves three
distinct chemical species: Substrate, Catalyst and
Link, hence \emph{SCL}. It was designed with a view
to generating simple phenomena of autopoietic
organisation. Varela et al.\fsp included a detailed
algorithmic account of their original model; however,
as documented in \cite{McMullin:OriginalAutop}, there
are a number of problems with interpreting and/or
re-implementing that algorithm. Arising from this,
that original algorithm was essentially set aside in
designing SCL; instead, SCL seeks to capture only the
general, qualitative, reaction schemes described by
Varela et al.
SCL was developed for two separate purposes.
Firstly, it provides a platform to critically
re-evaluate the phenomena---particularly autopoietic
phenomena---that can be realised with this general
kind of reaction scheme. That phenomenological
investigation will be described in a separate report.
The second objective was to gain experience of the
Swarm simulation system and evaluate its suitability
for this kind of project. This report is concerned
solely with this second objective: i.e.\fsp with
documenting the implementation, and with evaluating
the use of Swarm.
\vfill
\begin{quote}
\textbf{Keywords:} Autopoiesis, Artificial Life,
Artificial Chemistry, Swarm Simulation System.
\end{quote}
\end{abstract}
\begin{latexonly}
\tableofcontents
\end{latexonly}
\section{Introduction}
The qualitative inspiration for the chemistry
implemented by SCL is the model described in
\citeasnoun{Varela:Autopoiesis}. That model, including
the provided algorithm, has been reviewed in detail in
\cite{McMullin:OriginalAutop}, and will not be
presented again here. Instead, I will briefly describe
the qualitative chemical reaction scheme, and outline
the way SCL implements it. Note that the \emph{point}
of this particular reaction scheme is to model the
emergence and stability of \emph{autopoietic
organisation}; the reaction scheme was expressly
designed by Varela et al.\fsp with the specific idea
that it would be the simplest possible scheme that
could exhibit such organisation. However, the present
report is \emph{not} concerned with discussing or
elaborating the phenomenology actually generated by the
SCL model (autopoietic or otherwise)---that is deferred
for a subsequent, separate, report.
\section{Overview of the SCL Reaction Scheme}
SCL involves three distinct chemical species
(implemented as separate \emph{classes}):
\bigskip
\begin{tabular}{ll}
Substrate: & \sbst \\
Catalyst: & \cat \\
Link: & \lnk
\end{tabular}
\bigskip
The SCL reaction scheme is implemented in a 2-D space.
Subject to certain constraints, particles move around
this space in random walks. The particles can undergo
various transformations or reactions; where these
involve more than one particle, the reacting particles
must be adjacent to each other in the space.
The restriction to a 2-D space is important because it
means that linear chains or polymers of \lnk{} particles
can actually serve as spatial separators or boundaries.
It is stipulated that such chains are permeable to
\sbst{} particles, but impermeable to all others. In
this way a \emph{closed} chain of \lnk{} particles can
``trap'' \cat{} and \lnk{} particles inside, while still
allowing \sbst{} particles to move in (and out). These
constraints on particle motion are critical to the
intended autopoietic phenomena.
SCL supports six distinct reactions, as follows:
\begin{enumerate}
\item \emph{Production}:
\[\cat + 2 \sbst \rightarrow \cat + \lnk \]
In essence, this is a reaction in which two particles
of \sbst{} combine to produce one particle of \lnk.
However, the reaction can occur only with the mediation
of catalyst, \cat. The catalyst particle itself is
unaffected by the reaction (hence its name).\par
\item \emph{Disintegration}:
\[ \lnk \rightarrow 2 \sbst \]
This is a reverse reaction to production. It occurs
spontaneously.\par
\item \emph{Bonding}: Adjacent \lnk{} particles can
bond or polymerise into indefinitely long chains.
Each \lnk{} particle can form at most two bonds, so
that no structures more complex than chains can
arise. This is a spontaneous reaction, but it is
influenced in a complex way by the precise
configuration of \lnk{} particles involved, and by a
number of distinct parameters.
\lnk{} chains can break down again either via
spontaneous decay of individual bonds (see below), or
if constituent \lnk{} particles spontaneously
disintegrate back to substrate (when any associated
bonds will also undergo forced decay).\par
\item \emph{Bond Decay}: This is a reverse reaction to
bonding, in which individual bonds can decay,
breaking a chain. It occurs
spontaneously.\par
\item \emph{Absorption}:
\[ \lnk + \sbst \rightarrow \lnkplus \]
In this reaction, an \lnk{} particle can spontaneously
\emph{absorb} an \sbst{} particle. The absorbed \sbst{}
particle can subsequently be \emph{emitted} again (see
below). An \lnk{} which has absorbed an \sbst{} particle is
denoted by \lnkplus. An \lnk{} particle can absorb at
most one \sbst{} particle---i.e.\fsp an \lnkplus{} particle
cannot undergo a further absorption reaction. The
absorption and emission reactions together provide for
the permeability of \lnk{} chains to \sbst{} particles.\par
\item \emph{Emission}:
\[ \lnkplus \rightarrow \lnk + \sbst \]
This is a reverse reaction to absorption, in which a
previously absorbed \sbst{} particle is spontaneously
re-emitted. An \lnkplus{} particle which undergoes
disintegration must necessarily first undergo a
(forced) emission reaction.\par
\end{enumerate}
Each of these reactions is controlled by a one or more,
user configurable, rate parameters. Similarly, rates
of random motion in the space are controlled by
separate mobility parameters for each particle class.
This parameters are all implemented as class variables
of the relevant agent classes.
Of the six reactions, only the first three were present
in the original model described by Varela et al.
Furthermore, only disintegration was explicitly
controlled by a rate parameter, and no explicit mobility
parameters were defined, though there were implicit
differences in mobility between the different particle
classes. The \emph{implementation} of bonding in SCL is quite
different from the original.
Bond decay was added to SCL in the interests of making the
reaction scheme somewhat more symmetrical. However, if
desired, it can be ``switched off'' simply by setting
its rate parameter to zero.
The original model implemented permeability of \lnk{}
chains as a special case of \emph{motion} in the
space, rather than using the explicit absorption and
emission reactions introduced in SCL. However, that
suffered from the drawbacks of requiring a neighborhood
(for motion at least) extending two cells from
any given cell, and also making it more difficult to
control the rate at which permeation could occur
\emph{separately} from general mobility rates. I
suggest that introducing the explicit absorption and
emission reactions makes the model easier to understand
and provides more intuitively tractable parameters.
\section{Overview of the SCL Implementation}
SCL is implemented using the Swarm Simulation
System\footnote{\hoturl{http://www.santafe.edu/projects/swarm}}.
That is to say, SCL is written in Objective-C, using
the libraries supplied by Swarm, and is intended to be
portable across the computing platforms supported by
Swarm (Unix, running X-window and the \texttt{gcc}
compiler).
In what follows, it is assumed that the reader has a
basic understanding both of Objective-C and Swarm.
Unless explicitly stated otherwise, all references to
``random'' selection here imply selection with a
\emph{uniform} probability function over the relevant set.
\subsection{Space and Time}
SCL implements the reaction scheme in a discrete, 2D,
space with toroidal topology. Time is also discrete.
The space is organised as a square lattice. The
dynamics occur based on either the Moore (9-cell) or
the von Neumann (5-cell) neighborhood (this is, to some
extent, configurable). Each cell in the space is
occupied by a particle object. There are 4 subclasses
of particle objects, allowing for each of the three
distinct chemical species and a ``hole'' particle
class (denoted \hole). Bonds are also represented by
objects of another separate class. Lists are maintained
of all particles of each class, and of all bonds. The
basic execution cycle of the model, for one timestep,
is that each such list is traversed in turn, and each
particle or bond is sent a \texttt{step} message. The
effect of this message is, of course, class specific,
but generally involves possible motion (by swapping
position with an adjacent particle) followed by one or
more possible reactions.
In the following more detailed descriptions, the
particle which is actually executing its \texttt{step}
method will be called the \emph{primary} particle, to
distinguish it from the neighboring, or
\emph{secondary}, particle(s) which it may interact
with.
\subsection{Motion}
Motion for all particle classes follows the same
general strategy. In essence, particles undergo random
walks in the space. The relative rates of movement for
the different particle classes are controlled by user
configurable parameters.
Note that, on any given timestep, each particle gets
exactly one opportunity to be the primary particle of a
movement; but it will also get several (actually four)
opportunities to act as the secondary particle of a
movement.
In more detail, movement takes place as follows:
\begin{enumerate}
\item Firstly, particles may, at any given time, be
either mobile or immobile. The exact circumstances
governing this are class specific. However, for all
classes, once a particle moves, it then becomes
immobile for the remainder of that timestep. That
is, no particle may move more than once within a single
timestep.\par
\item Assuming a given primary particle \emph{is} mobile, a
neighboring cell is chosen at random from the von
Neumann neighborhood. The particle in this cell is
then the \emph{secondary} particle---a candidate for
swapping with the target particle.\footnote{It
follows from this choice of neighborhood that each
particle gets exactly four opportunities to be
considered as the secondary particle of a movement,
per timestep. The von Neumann neighborhood is used
here because this makes implementation of
\emph{im}-permeability of the \lnk{} particle chains
relatively simple.} The mobility status of this
candidate particle is checked.\par
\item Assuming that the neighboring particle \emph{is}
also mobile, a bernoulli random variable (biased coin
toss) is evaluated to determine whether to proceed
with the swap or not. The probability parameter
used in this evaluation thus allows control of the
relative rates of movement.
In an earlier implementation of SCL, this probability
of movement was simply a parameter of the class of
the primary particle. However, this is not
completely satisfactory because a particle may move
as a result of being a primary particle or as a
secondary particle---and, on that original scheme,
the probabilities in the two cases, for the same two
particles, could be different, which is somewhat
counter-intuitive.
In SCL v0.04 each particle class still has a class
specific mobility parameter, called the
\texttt{MobilityFactor}. But rather than simply
taking the probability to be the
\texttt{MobilityFactor} of the primary particle
class, it is calculated as the geometric mean of the
\texttt{MobilityFactor} parameters for the classes of
the primary and secondary particles. This means that
the probability is always the same for swapping
particles of any two particular classes, regardless
of which is primary and which is secondary. Further,
by using the geometric (rather than, say, arithmetic)
mean, we ensure that if the \texttt{MobilityFactor}
is set to \emph{zero} for either class, then the
overall probability is guaranteed to be zero.
Finally, of course, if the \texttt{MobilityFactor}
for both classes is the same, then this just
\emph{is} the probability of the particular
swap.
Note that the primary reason for introducing the
\hole{} particle class was so that this movement
strategy could be applied uniformly. The alternative
would be that holes in the space would be represented
by positions with no object present at all (i.e.\fsp the
Objective-C \texttt{nil} object pointer)---but then
there would be no corresponding
\texttt{MobilityFactor} and nothing to ``swap'' with
either. While using explicit \hole{} particles
instead presumably involves some performance
penalty, I considered this worthwhile in order to
keep the implementation simpler and more
consistent.\par
\item Assuming that the bernoulli random variable evaluates as
\texttt{YES}, the two particles swap position. Both
then become immobile for the remainder of that timestep.
\end{enumerate}
\emph{\lnk{} particles with one or more bonds are always
immobile.} This is a very strong restriction on
the model. It was present in the original model of
Varela et al.\fsp and is retained here. The presumed
reason for this restriction is to avoid the
rather complicated issue of what should happen to a bond
(and the \lnk{} particle at its other end) if a bonded
\lnk{} particle is allowed to move.
\subsection{Reactions}
The reactions are all initiated and directed by the
execution of the \texttt{step} method on some
particular primary particle or on a bond. Which
reactions are considered is determined by the particle
class. The production reaction can be initiated only
by \cat{} particles; the bond decay reaction is
initiated by the bond itself; all other reactions are
initiated only by \lnk{} particles.
Reactions typically occur in either the von Neumann
or Moore neighborhood of a given primary particle. A
user configurable parameter determines a single
\emph{default neighborhood} which applies to all such
reactions.
\subsubsection{Production}
Production is initiated by a primary \cat{} particle,
and is implemented as follows:
\begin{enumerate}
\item A bernoulli random variable is evaluated with
probability parameter \texttt{productionProbability}
(the rate parameter for the reaction). If this
evaluates as \texttt{NO}, the reaction terminates.\par
\item A random neighbor is chosen from the default
neighborhood (\texttt{neighborParticle0}). If this is
\emph{not} an \sbst{} particle, the reaction is
terminated.\par
\item There will be exactly two cells in the
default neighborhood which are immediately adjacent
to both the primary \cat{} particle and the
\texttt{neighborParticle0} \sbst{} particle. One of
these is chosen at random (\texttt{neigborParticle1}).
If this does \emph{not} contain an \sbst{} particle, the
reaction is terminated.\par
\item \texttt{neighborParticle0} is replaced with an
\hole; \texttt{neighborParticle1} is replaced with an
\lnk. That completes the reaction.
\end{enumerate}
\subsubsection{Disintegration}
Disintegration is implemented by a primary \lnk{}
particle, and happens in two quite separate stages.
Each \lnk{} particle has an associated boolean state
variable called \texttt{disintegrating}. When an \lnk{}
particle is initially created (via production) this is
set to \texttt{NO}. The first phase of disintegration
is for this state variable to change to \texttt{YES},
and happens as follows:
\begin{enumerate}
\item A bernoulli random variable is evaluated with
probability parameter \texttt{disintegrationProbability}
(the rate parameter for the reaction). If this
evaluates as \texttt{NO}, the reaction terminates.\par
\item The \texttt{disintegrating} state variable is set
to \texttt{YES}. This completes the first stage of
the reaction.
\end{enumerate}
Once an \lnk{} particle goes into the disintegrating
state it stays in that state until the disintegration
can be completed. In general it cannot be completed
immediately because it relies on finding one or more
adjacent holes to accommodate the disintegration
products.
An \lnk{} particle in the disintegrating state attempts
to complete the reaction as follows (this attempt
occurs for the first time on the \emph{same} timestep
as the \lnk{} particle goes into the disintegrating
state; it will be repeated, as necessary, on each
following timestep, until the reaction is completed):
\begin{enumerate}
\item If the \lnk{} particle has an absorbed \sbst{}
particle (i.e. it is strictly an \lnkplus), then an
attempt is made to forcibly emit this \sbst. The
details of this process are given \hyperref{below}{in
section~}{}{emission}.\par
\item If the \lnk{} particle still has an absorbed
\sbst{} particle (i.e.\fsp the attempted emission failed)
then the reaction is terminated.\par
\item A random neighbor is chosen from the default
neighborhood (\texttt{neighborParticle}). If this is
\emph{not} a \hole, the reaction is terminated.\par
\item The \hole{} is replaced with an \sbst{} particle.\par
\item Any bonds associated with the primary \lnk{}
particle are forced to decay (see
\hyperref{below}{section~}{}{bonddecay}).\par
\item The primary \lnk{} particle is replaced with an
\sbst{} particle. This completes the disintegration
reaction.
\end{enumerate}
Note finally that once an \lnk{} particle has gone into
the disintegrating state it will no longer undergo the
absorption reaction (since this could further delay the
completion of the disintegration); however, it can
still participate in bonding.
\subsubsection{Bonding}
Bonding is initiated by a primary \lnk{} particle;
however, in general, the various constraints or
influences affecting bonding are evaluated
symmetrically for both the primary and secondary \lnk{}
particle.
The factors influencing the bonding reaction
are as follows:
\begin{itemize}
\item Bonding is inhibited if an \lnk{} particle has
already formed two bonds. This sets the maximum
number of bonds to any \lnk{} particle as two, and
ensures that the only particle aggregates or
``molecules'' which can form are linear ``chains''
rather than any more complex, branching, structures.\par
\item Two other mechanisms or interactions by which
bonding can be inhibited are also provided. Each of
these is qualified by a \texttt{BOOL} parameter
whereby they can be enabled or disabled. These are
essentially intended as alternative mechanisms for
preventing ``premature'' bonding of free \lnk{}
particles that have been produced within the boundary
of an autopoietic entity. However, even though the
typical mode of operation would be with just one or
the other of these interactions enabled, SCL will
actually allow both to be simultaneously disabled, or
both simultaneously enabled, if desired.
The need for \emph{some} such form of inhibition
interaction is quite subtle but, it transpires, quite
crucial to the autopoietic phenomenology of the
model; while no such effect was mentioned explicitly
in \cite{Varela:Autopoiesis}, it is almost certain
that it was present in their model. This is discussed
further in \cite{McMullin:OriginalAutop}.
\begin{itemize}
\item \emph{Inhibition by a neighboring \lnk{} Chain}: This
interaction allows bonding to be inhibited to any
\lnk{} particle which is \emph{alongside} (as opposed
to, at the end of) an existing chain of
\lnk{} particles. This is implemented for any given
\lnk{} particle by checking whether there are any existing
doubly bonded \lnk{} particles in its (Moore)
neighborhood. This interaction is qualified by the
\texttt{BOOL} parameter, \texttt{chainInhibitBondFlag}.
If this is \texttt{YES} this inhibition effect is
enabled; if it is \texttt{NO} then it is
disabled.
As discussed in \cite{McMullin:OriginalAutop}, this
particular form of bond inhibition reaction was
inspired by examination of the \texttt{FORTRAN IV}
code for one early version of the model developed by
Varela et al.\par
\item \emph{Inhibition by a neighboring \cat{}
particle}: This is implemented for any given \lnk{}
particle by checking whether there are any \cat{}
particles in its (Moore) neighborhood. This
interaction is qualified by the \texttt{BOOL}
parameter, \texttt{catInhibitBondFlag}. If this is
\texttt{YES} this inhibition effect is enabled; if it
is \texttt{NO} then it is disabled.
This alternate form of bond inhibition reaction was
inspired by \cite{Lizana:Autopoiesis}.
\end{itemize}\par
\item A second bond to any particular \lnk{} particle
must form an angle of $90^\circ$ or more relative to
the first bond. This rules out forming two bonds ``on
top of each other'' (a double bond between the same
two \lnk{} particles), or with an angle of just
$45^\circ$ between them.\par
\item Given that a new bond would satisfy the above
constraints for both affected \lnk{} particles, the
formation of the bond is controlled by a reaction
rate, or probability parameter. More precisely, one
of three possible rate parameters is applied,
depending on the exact conformation of the proposed
new bond:
\begin{enumerate}
\item If neither \lnk{} particle already has a bond, the
new bond would initiate a new chain; in that case the
probability is set by the parameter
\texttt{chainInitiateProbability}.
\item If just one of the two \lnk{} particles already
has a bond, then the new bond would extend an already
existing chain by one more particle; in that case the
probability is set by the parameter
\texttt{chainExtendProbability}.
\item If both of the \lnk{} particles already have a
bond, then the new bond would splice together two
existing chains; in that case the
probability is set by the parameter
\texttt{chainSpliceProbability}.
\end{enumerate}
\end{itemize}
The detailed implementation of the bonding reaction
is as follows:
\begin{enumerate}
\item If bonding to the primary \lnk{} is inhibited due
to already having two bonds, or either of the
inhibition interactions (an adjacent doubly bonded
\lnk{} particle, qualified by
\texttt{chainInhibitBondFlag}, or an adjacent \cat{}
particle, qualified by \texttt{catInhibitBondFlag})
the reaction is terminated.\par
\item A random neighbor (\texttt{neighborParticle}) is
chosen from the default neighborhood.\par
\item If the primary \lnk{} already has a bond to this
neighbor, or to either of the neighbors at an angle of
$45^\circ$ to this neighbor, the reaction is
terminated.\par
\item If \texttt{neighborParticle} is not an
\lnk{} particle the reaction is terminated. Otherwise
it becomes the secondary \lnk{} particle in the reaction.\par
\item If bonding to the secondary \lnk{} is inhibited
due to already having two bonds, or either of the
inhibition interactions, the reaction is
terminated.\par
\item If the secondary \lnk{} already has a bond to
either of its neighbors at an angle of $45^\circ$ to
the direction from it to the primary \lnk{} particle, the
reaction is terminated.\par
\item If neither the primary nor secondary
\lnk{} particles already have a bond,
\texttt{bondingProbability} will be set to
\texttt{chainInitiateProbability}; if just one of them
already has a bond, \texttt{bondingProbability} will be set to
\texttt{chainExtendProbability}; if both of them already
have a bond, \texttt{bondingProbability} will be set to
\texttt{chainSpliceProbability}.\par
\item A bernoulli random variable is evaluated with
probability parameter \texttt{bondingProbability};
if this evaluates as \texttt{NO}, the
reaction terminates.\par
\item The secondary \lnk{} particle creates the bond to
the primary \lnk{} particle (and passes a pointer to
it back to the latter).\par
\item The primary \lnk{} particle records the presence
of the new bond also. This completes the bonding
reaction.
\end{enumerate}
\subsubsection{Bond Decay\label{bonddecay}}
Bond decay can happen either as a spontaneous reaction,
or as a forced reaction when an \lnk{} is completing the
disintegration reaction. The spontaneous reaction is
initiated by a bond, and is implemented as follows:
\begin{enumerate}
\item A bernoulli random variable is evaluated with
probability parameter \texttt{bondDecayProbability}
(the rate parameter for the reaction). If this
evaluates as \texttt{NO}, the reaction terminates.\par
\item The bond decays (i.e.\fsp the two affected \lnk{}
particles are told to remove the bond). This
completes the spontaneous bond decay reaction.
\end{enumerate}
\newpage
The forced bond decay reaction is initiated by one of
the bonded \lnk{} particles, as it completes the
disintegration reaction. It is implemented as follows:
\begin{enumerate}
\item The primary \lnk{} particle tells the bond to
decay.
\item The bond decays (i.e.\fsp the two affected \lnk{}
particles are told to remove the bond). This
completes the forced bond decay reaction.
\end{enumerate}
\subsubsection{Absorption}
Absorption is initiated by a primary \lnk{} particle,
and is implemented as follows:
\begin{enumerate}
\item If the primary \lnk{} particle already has an
absorbed link (i.e.\fsp it is strictly an \lnkplus{}
particle) then the reaction terminates.
\item A bernoulli random variable is evaluated with
probability parameter \texttt{absorptionProbability}
(the rate parameter for the reaction). If this
evaluates as \texttt{NO}, the reaction terminates.\par
\item A random neighbor is chosen from the default
neighborhood (\texttt{neighborParticle}). If this is
\emph{not} an \sbst{} particle, the reaction
terminates.\par
\item \texttt{neighborParticle} is replaced with a
hole.
\item The primary \lnk{} particle is marked as having an
absorbed substrate---i.e.\fsp it changes to the
\lnkplus{} form. That completes the absorption
reaction.
\end{enumerate}
\subsubsection{Emission\label{emission}}
Emission is initiated by a primary \lnk{} particle. It
can happen either as a spontaneous reaction, or as a
forced reaction when an \lnk{} is in the disintegrating state.
The spontaneous reaction is implemented as follows:
\begin{enumerate}
\item If the primary \lnk{} particle does not have an
absorbed link (i.e.\fsp it is not in the \lnkplus{} form)
then the reaction terminates.
\item A bernoulli random variable is evaluated with
probability parameter \texttt{emissionProbability}
(the rate parameter for the reaction). If this
evaluates as \texttt{NO}, the reaction terminates.\par
\item A random neighbor is chosen from the default
neighborhood (\texttt{neighborParticle}). If this is
\emph{not} a hole (\hole), the reaction
terminates.\par
\item \texttt{neighborParticle} is replaced with an
\sbst{} particle.
\item The primary \lnkplus{} particle is marked as no
longer having an absorbed substrate---i.e.\fsp it
reverts to the \lnk{} form. That completes the
absorption reaction.
\end{enumerate}
In the case where the emission reaction is forced, as a
side effect of the disintegration reaction, the
implementation is the same as this spontaneous reaction
except that steps~1 and~2 above are bypassed.
\section{Getting SCL}
SCL is available only as a source archive. See
\hyperref{the Retrieval section}
{section~}{}{retrieval} for information on retrieving
it. Unpack the archive file into an empty
directory. Edit the \texttt{SWARMHOME} variable in
\texttt{Makefile} as appropriate,
and type \texttt{make}. Provided Swarm itself has been
correctly installed on your system (and is a compatible
release) then SCL should now be built.
SCL v0.04 was originally developed against the Swarm
release dated \texttt{961002}; since this is still a
\emph{beta} release, later releases will not
necessarily be backward compatible, and SCL may well
require modification to successfully build against any
such later Swarm release.
\section{Using SCL}
When SCL starts up it initially creates three windows to
facilitate user interaction:
\begin{itemize}
\item \texttt{Swarm}
\item \texttt{ParameterManager}
\item \texttt{WorldManager}
\end{itemize}
The latter two of these are Swarm \texttt{probeDisplay}
windows on specifically designed user interaction
objects; the standard Swarm documentation provides
general information on the facilities and interactions
offered by \texttt{probeDisplay} windows, and that will
not be repeated here. These initial three windows are
required throughout a session with SCL, and none should
be deleted by the user.
Once a model world is created or loaded from disk, two
additional windows are created by SCL:
\begin{itemize}
\item \texttt{SCL World}
\item \texttt{UserCellEditor}
\end{itemize}
All of these user interaction windows are discussed in
detail in the following sections.
\subsection{\texttt{SWARM} Window}
\begin{rawhtml}
 \end{rawhtml}
This is the standard Swarm \emph{control panel} window.
It provides buttons to start the simulation
(\texttt{Go}), stop it (\texttt{Stop}), single step
(\texttt{Timestep}) and terminate the simulation
(\texttt{Quit}). The buttons for controlling execution
of the simulation will, of course, operate only while a
simulation model, or world, exists. The tools for
creating and manipulating these world models will be
described separately below. This window is \emph{not}
generated as a \texttt{probeDisplay} on some object,
but is rather generated automatically as a standard
feature of Swarm.
\begin{rawhtml}
\end{rawhtml}
This is the standard Swarm \emph{control panel} window.
It provides buttons to start the simulation
(\texttt{Go}), stop it (\texttt{Stop}), single step
(\texttt{Timestep}) and terminate the simulation
(\texttt{Quit}). The buttons for controlling execution
of the simulation will, of course, operate only while a
simulation model, or world, exists. The tools for
creating and manipulating these world models will be
described separately below. This window is \emph{not}
generated as a \texttt{probeDisplay} on some object,
but is rather generated automatically as a standard
feature of Swarm.
\begin{rawhtml}
\end{rawhtml}
\subsection{\texttt{ParameterManager} Window}
\begin{rawhtml}
 \end{rawhtml}
This is a (tailored) \texttt{probeDisplay} on the
\texttt{parameterManager} object. It provides
facilities to monitor and modify all the model
parameters supported by SCL, and for saving and/or
reloading these parameters to or from disk files.
To modify a parameter, simply click on the relevant
position in the window and edit the value. Ensure that
you press \texttt{Enter} or \texttt{Return} to cause
the new value to take effect (this is a standard
feature of Swarm \texttt{probeDisplay} windows). Note
that, because the \texttt{probeDisplay} is continuously
refreshed while the simulation is running, it is only
practical to alter parameters while the simulation is
stopped.
Note also that no attempt has been made to make the
file interface ``robust''; specifically, any error in
accessing a file (e.g.\fsp if it does not exist and/or
cannot be created, or is of the wrong format etc.)
will result in a fatal exception being raised and SCL
will exit.
\begin{rawhtml}
\end{rawhtml}
This is a (tailored) \texttt{probeDisplay} on the
\texttt{parameterManager} object. It provides
facilities to monitor and modify all the model
parameters supported by SCL, and for saving and/or
reloading these parameters to or from disk files.
To modify a parameter, simply click on the relevant
position in the window and edit the value. Ensure that
you press \texttt{Enter} or \texttt{Return} to cause
the new value to take effect (this is a standard
feature of Swarm \texttt{probeDisplay} windows). Note
that, because the \texttt{probeDisplay} is continuously
refreshed while the simulation is running, it is only
practical to alter parameters while the simulation is
stopped.
Note also that no attempt has been made to make the
file interface ``robust''; specifically, any error in
accessing a file (e.g.\fsp if it does not exist and/or
cannot be created, or is of the wrong format etc.)
will result in a fatal exception being raised and SCL
will exit.
\begin{rawhtml}
\end{rawhtml}
\subsection{\texttt{WorldManager} Window}
\begin{rawhtml}
 \end{rawhtml}
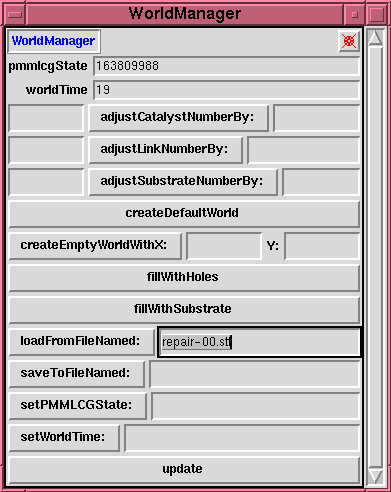
This is a (tailored) \texttt{probeDisplay} on the
\texttt{worldManager} object. It provides ``high
level'' facilities for creating and manipulating a
model world (or, more precisely, a model world
\emph{state}), and for saving and/or reloading the
world state to or from disk files.
Note that if the world state is loaded from disk then
any already loaded world model will be automatically
(and silently) dropped first. As with the file
interface for parameters, no attempt has been made to
make the world state file interface ``robust'', and any
error in accessing a file will result in a fatal
exception being raised and SCL will exit.
\texttt{PMMLCG} refers to the pseudo random number
generator object used to implement the (pseudo!)
stochastic effects in SCL. The \texttt{WorldManager}
window allows the exact state of this generator to be
monitored and/or modified. This permits, for example,
a given world state to be reloaded and then restarted
with a different initial value for the generator state,
thus potentially exploring the robustness of stochastic
phenomena in the model.
The \texttt{worldTime} records the number of full
timesteps which have elapsed. It is meaningful only
when a world model has been created or loaded from
disk.
The \texttt{WorldManager} window offers two ways of
creating a new model world. As with loading a model
from disk, if a new model is created here then any
already loaded model will be automatically (and
silently) dropped first. The
\texttt{createDefaultWorld} method provides a
``canned'' world with certain compiled-in
characteristics. The \texttt{createEmptyWorldWithX:Y:}
method allows a world of arbitrary size to be created.
This will be initially ``empty'', i.e.\fsp populated
exclusively with holes.
Once a world has been created (or loaded from disk)
particles can be incrementally added to (or removed
from) the world using the three
\texttt{adjust...NumberBy:} methods---these accept a
positive or negative integer argument, being the number
of particles to add or remove respectively. Particles
are added or removed at random positions with these
methods. They can be added only into holes; when they
are removed, they leave holes. These methods return the
number of particles \emph{actually} added or removed.
This may generally differ from the number requested.
The methods can only add new particles into existing
holes---so if the world becomes full, no more can be
added. Conversely, only existing particles can be
removed---so once all particles of a given class have
been removed no more can be taken out.
Special methods are also provided to forcibly fill the
space completely with substrate, or completely empty it
again (i.e.\fsp ``fill'' with holes).
\begin{rawhtml}
\end{rawhtml}
This is a (tailored) \texttt{probeDisplay} on the
\texttt{worldManager} object. It provides ``high
level'' facilities for creating and manipulating a
model world (or, more precisely, a model world
\emph{state}), and for saving and/or reloading the
world state to or from disk files.
Note that if the world state is loaded from disk then
any already loaded world model will be automatically
(and silently) dropped first. As with the file
interface for parameters, no attempt has been made to
make the world state file interface ``robust'', and any
error in accessing a file will result in a fatal
exception being raised and SCL will exit.
\texttt{PMMLCG} refers to the pseudo random number
generator object used to implement the (pseudo!)
stochastic effects in SCL. The \texttt{WorldManager}
window allows the exact state of this generator to be
monitored and/or modified. This permits, for example,
a given world state to be reloaded and then restarted
with a different initial value for the generator state,
thus potentially exploring the robustness of stochastic
phenomena in the model.
The \texttt{worldTime} records the number of full
timesteps which have elapsed. It is meaningful only
when a world model has been created or loaded from
disk.
The \texttt{WorldManager} window offers two ways of
creating a new model world. As with loading a model
from disk, if a new model is created here then any
already loaded model will be automatically (and
silently) dropped first. The
\texttt{createDefaultWorld} method provides a
``canned'' world with certain compiled-in
characteristics. The \texttt{createEmptyWorldWithX:Y:}
method allows a world of arbitrary size to be created.
This will be initially ``empty'', i.e.\fsp populated
exclusively with holes.
Once a world has been created (or loaded from disk)
particles can be incrementally added to (or removed
from) the world using the three
\texttt{adjust...NumberBy:} methods---these accept a
positive or negative integer argument, being the number
of particles to add or remove respectively. Particles
are added or removed at random positions with these
methods. They can be added only into holes; when they
are removed, they leave holes. These methods return the
number of particles \emph{actually} added or removed.
This may generally differ from the number requested.
The methods can only add new particles into existing
holes---so if the world becomes full, no more can be
added. Conversely, only existing particles can be
removed---so once all particles of a given class have
been removed no more can be taken out.
Special methods are also provided to forcibly fill the
space completely with substrate, or completely empty it
again (i.e.\fsp ``fill'' with holes).
\begin{rawhtml}
\end{rawhtml}
\subsection{\texttt{SCL World} Window}
\begin{rawhtml}
 \end{rawhtml}
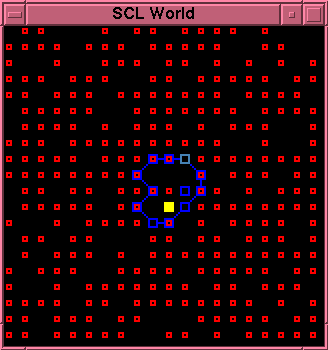
This is a graphic image of the state of the model.
This is implemented with the Swarm \texttt{ZoomRaster}
class, and so can be magnified or reduced (by dragging
any corner). Note however that this does \emph{not}
allow ``windowing'' onto a subarea of the world model,
or panning across it: this window will always display
the entire world. If this window is resized while the
model is stopped the image will \emph{not} be
automatically updated; you can use the
\texttt{update} button in the
\texttt{WorldManager} probe to force an update.
\newpage \sbst{} particles are shown as small,
outlined, red squares; \cat{} particles are shown as
larger, filled, yellow squares. Bonds are blue line
segments. \lnk{} particles are normally shown as blue,
outlined, squares, the same size as \cat{} particles.
If an \lnk{} particle goes into the disintegrating
state then it turns grey. If an \lnk{} particle has an
absorbed substrate (i.e.\fsp it is in the \lnkplus{}
form) then this absorbed particle is still shown as a
small, outlined, red square, but internal to the \lnk{}
particle.
Moving the pointer over this window and clicking mouse
button~1 (normally the left button) will set the
co-ordinates of the \texttt{UserCellEditor}---which is
discussed in section~\ref{celleditor} below.
Moving the pointer over this window and clicking mouse
button~3 (normally the right button) will create a new
\texttt{probeDisplay} for the particle object pointed
at. These particle objects will not be detailed
further here---consult the source code to see exactly
what instance variables and methods are associated with
any particular particle class. However, it is worth
noting that this is the \emph{only} mechanism for a
user to manipulate or modify \emph{bonds}---i.e.\fsp by
creating a \texttt{probeDisplay} on an affected \lnk{}
particle, and using the relevant methods provided
there. There is \emph{no} mechanism to create a
\texttt{probeDisplay} for a bond object.
Be aware that, due to an underlying problem with the
current release of Swarm, SCL will crash if any object,
which is being probed with a \texttt{probeDisplay}, is
dropped. Since particle objects in Swarm may quite
generally get dropped, it is recommended that you only
a create \texttt{probeDisplay} on a particle object
while the simulation is stopped; and that you drop this
\texttt{probeDisplay} (using the special, red, destroy
button---\emph{not} your window manager) before
continuing execution of the model.
\begin{rawhtml}
\end{rawhtml}
This is a graphic image of the state of the model.
This is implemented with the Swarm \texttt{ZoomRaster}
class, and so can be magnified or reduced (by dragging
any corner). Note however that this does \emph{not}
allow ``windowing'' onto a subarea of the world model,
or panning across it: this window will always display
the entire world. If this window is resized while the
model is stopped the image will \emph{not} be
automatically updated; you can use the
\texttt{update} button in the
\texttt{WorldManager} probe to force an update.
\newpage \sbst{} particles are shown as small,
outlined, red squares; \cat{} particles are shown as
larger, filled, yellow squares. Bonds are blue line
segments. \lnk{} particles are normally shown as blue,
outlined, squares, the same size as \cat{} particles.
If an \lnk{} particle goes into the disintegrating
state then it turns grey. If an \lnk{} particle has an
absorbed substrate (i.e.\fsp it is in the \lnkplus{}
form) then this absorbed particle is still shown as a
small, outlined, red square, but internal to the \lnk{}
particle.
Moving the pointer over this window and clicking mouse
button~1 (normally the left button) will set the
co-ordinates of the \texttt{UserCellEditor}---which is
discussed in section~\ref{celleditor} below.
Moving the pointer over this window and clicking mouse
button~3 (normally the right button) will create a new
\texttt{probeDisplay} for the particle object pointed
at. These particle objects will not be detailed
further here---consult the source code to see exactly
what instance variables and methods are associated with
any particular particle class. However, it is worth
noting that this is the \emph{only} mechanism for a
user to manipulate or modify \emph{bonds}---i.e.\fsp by
creating a \texttt{probeDisplay} on an affected \lnk{}
particle, and using the relevant methods provided
there. There is \emph{no} mechanism to create a
\texttt{probeDisplay} for a bond object.
Be aware that, due to an underlying problem with the
current release of Swarm, SCL will crash if any object,
which is being probed with a \texttt{probeDisplay}, is
dropped. Since particle objects in Swarm may quite
generally get dropped, it is recommended that you only
a create \texttt{probeDisplay} on a particle object
while the simulation is stopped; and that you drop this
\texttt{probeDisplay} (using the special, red, destroy
button---\emph{not} your window manager) before
continuing execution of the model.
\begin{rawhtml}
\end{rawhtml}
\subsection{ \texttt{UserCellEditor} Window\label{celleditor}}
\begin{rawhtml}
 \end{rawhtml}
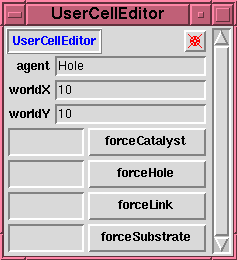
This is a (tailored) \texttt{probeDisplay} on the
\texttt{userCellEditor} object. It provides ``low
level'' facilities for manipulating the model world.
In particular it allows the user to ``force'' a
particle of any class at any cell in the world. It is
used by first shifting its coordinates to the desired
cell---by clicking mouse button 1 while pointing at the
cell, as mentioned above---and then forcing the desired
particle class at that cell. If you wish to modify the
particle in any further way (especially to create bonds
between \lnk{} particles) you must create a
\texttt{probeDisplay} on the particle itself and use
that, as already discussed.
\begin{rawhtml}
\end{rawhtml}
This is a (tailored) \texttt{probeDisplay} on the
\texttt{userCellEditor} object. It provides ``low
level'' facilities for manipulating the model world.
In particular it allows the user to ``force'' a
particle of any class at any cell in the world. It is
used by first shifting its coordinates to the desired
cell---by clicking mouse button 1 while pointing at the
cell, as mentioned above---and then forcing the desired
particle class at that cell. If you wish to modify the
particle in any further way (especially to create bonds
between \lnk{} particles) you must create a
\texttt{probeDisplay} on the particle itself and use
that, as already discussed.
\begin{rawhtml}
\end{rawhtml}
\section{SCL Design Notes}
SCL demonstrates possible approaches to some Swarm
programming issues that are not covered in the sample
applications distributed with the current
(\texttt{961002}) release of Swarm, and which may
therefore be of interest to the broader Swarm
community. The definitive reference here is, of
course, to the source code itself. But a brief
introduction will be given in the following sections.
\subsection{Who needs an Observer?}
The standard Swarm example applications enforce a strict
distinction between an ``observer'' and the ``model''
itself. The idea is to facilitate easier switching
between versions which operate in ``batch'' mode (with
no user interaction) and versions which operate in
``GUI'' mode (with user interaction).
This is a very reasonable and useful strategy.
However, in the case of SCL I knew that I did not want
the facility to operate in ``batch''mode---it was
intended and designed strictly as an interactive
application. Furthermore, I found that the attempt to
maintain the flexibility of supporting both batch and
interactive modes made the application more complex,
and also made the implementation of user interaction
more cumbersome. Therefore I abandoned the
observer/model distinction, and rationalised the
program accordingly. This is not an intrinsically
better or worse approach; it is simply another
alternative that some users may find of interest.
\subsection{Class Variables}
Objective C does not strictly offer ``class variables'',
or, at least, has no explicit support for them. However
the effect of such variables \emph{can} be realised by
using static variables with file scope in the class
implementation file. Note, however, that these
variables will \emph{not} be accessible to subclasses.
Class variables are useful in SCL to implement
reaction and mobility parameters. These must be common
to all objects of a specified class. They could be
implemented with conventional global variables; but
since they \emph{are} class specific, it is more
appropriate to implement them as class variables.
A problem with this approach is that the Swarm probe
machinery does not support probing of class objects.
Thus, probes cannot be used \emph{directly} to give
user access to these parameters. I understand that
this is a fairly fundamental limitation, and is
unlikely to be removed in the future.
However, this is easily worked around by introducing a
special object, \texttt{parameterManager} which
``shadows'' the class variables as instance variables.
This can then be probed as usual to provide access to
the parameters. Some care is required in using this
approach. One can normally use a probe to directly
modify instance variables; one can do that in this case
also, but it will not automatically have the desired or
expected effect, because such modifications do not, in
themselves, affect the class variables. To correct
this, the \texttt{parameterManager} provides the
\texttt{apply} method which essentially copies all the
instance variables into the corresponding class
variables. To make the behaviour as robust as
possible, an invocation of \texttt{apply} in included
in \emph{every} timestep---so that any changes in the
instance variables will automatically take full effect at
the start of the next timestep. Similarly,
\texttt{apply} is invoked immediately prior to saving a
model to disk.
\subsection{2D Space/2D Agents}
Swarm provides a standard class (\texttt{Discrete2d})
for implementing a discrete, 2-D, space, with an
arbitrary object (or \texttt{nil}) in each cell. This
class does not provide for any special treatment or
handling of the boundary of this space. A subclass of
this, \texttt{Grid2d} adds functionality to check that
a new object is not inserted in the space overwriting a
previous object. The sample \texttt{heatbugs}
application uses this class, and additionally imposes a
toroidal topology. The calculations for the latter are
carried out in the \texttt{heatbug} agents; because
these calculations are rather frequent, the size of the
space is cached in the agents.
In SCL I wanted both this toroidal topology, and also
the restriction to at most one agent (particle in my
case) in any single cell. However, the approach in SCL
is somewhat different---indeed, it takes an almost
diametrically opposed approach to where to locate these
functionalities.
First, instead of locating the calculations for the
toroidal topology in the agents, I developed a new
space class, \texttt{DiscreteToroid}, subclassed from
\texttt{Discrete2d}, which offers explicit methods for
adjusting (wrapping) co-ordinates appropriately. I
consider this preferable, because the space object is
the appropriate object to ``know'' how to deal with
co-ordinates that go outside the defined boundaries.
However, it does mean sacrificing some performance
compared to the \texttt{heatbugs} approach. In my
particular case, performance has not been a problem, so
this was a reasonable change; it would not necessarily
be so in other models.
Secondly, instead of having the space attempt to
restrict occupation of any cell to one object at a
time, I embedded this functionality in a new agent
class, \texttt{Agent2d}, which became the superclass
for all particle classes. \texttt{Agent2d} also
encapsulates a lot of other ``generic'' functionality
for agents in a 2D space---at least where interactions
are local (in the von Neumann or Moore neighborhoods),
and include movement by swapping. \texttt{Agent2d} may
thus be useful in other models, quite different from
SCL. \texttt{Agent2d} functionality is layered, in
part, on top of two other new agent classes,
\texttt{VNneighbor} and \texttt{MooreNeighbor} which
provide support for co-ordinate calculations within
these particular neighborhoods.
\subsection{Mortal Agents!}
It is in the nature of the SCL model that agents
(particles) are created and destroyed on an on-going
basis during the simulation. This led to two separate
problems.
Firstly, there is a general problem with Swarm that the
application will crash if any object which is being
probed (with a \texttt{probeDisplay}) is dropped. I
have not attempted to provide any fix for this. It can
be worked around, as already explained, by taking care
to ensure that \texttt{probeDisplay} windows on particles
are created only while the simulation is stopped, and
are dropped before the simulation is continued. It
would probably be possible to \emph{enforce} this
restriction from within the program; but, since this
will probably be fixed ``properly'' in a future Swarm
release, I did not consider that the effort was
justified.
Secondly---and more seriously---the scheduling of model
execution can be disrupted, leading to a crash, if
agents are dropped during the simulation. This arises
where a list of agents is being traversed (via a
\texttt{ForEach:} method), and, within that traversal,
an object is removed from the list. This is a
consequence of the way lists are currently implemented
in Swarm. Again, the restriction should be removed in
a future Swarm release. However, since this behaviour
was essential to SCL it was absolutely necessary to
develop a workaround.
The strategy adopted is to have a ``manager'' object
for the agents. This accepts requests to create or
drop agents and effectively puts these requests into a
\emph{queue}. A certain amount of processing of the
request can be (indeed, must be) done immediately; but
the list of agents which is potentially being traversed
by \texttt{ForEach:} is not immediately updated.
Instead, that final update is deferred until some time
later, when it is known to be safe.
This mechanism is implemented in SCL via a generic
\texttt{AgentManager} class; this is separately
instantiated to handle each of the different classes of
agent required (one for bonds, and one each for the
different particle classes, including holes).
This mechanism works, but is quite cumbersome and
somewhat error prone (care is required in ensuring that
the various queues are processed when necessary but
also when safe). It should probably be dispensed with
as soon as Swarm can handle agent dropping safely. On
the other hand, this mechanism does offer an
interesting potential (\emph{not} currently exploited)
for improving performance. The \texttt{AgentManager}
could quite easily arrange to \emph{cache} dropped
agents, and recycle them in response to new creation
requests. This might significantly reduce the overhead
associated with creating and/or dropping agents.
However, some care would be required to ensure that
this ``pseudo'' creation was equivalent to ``genuine''
creation. This latter issue is potentially complicated
by Swarm's powerful, but complex, ``create phase
protocol''.
\subsection{Agent Update Order}
SCL implements a single timestep by processing each
agent list in turn, issuing a \texttt{step} message to
each agent on that list. The order in which the
different agent lists (or classes) are processed is
fixed. \emph{Within} each agent list, processing is
done in the list's ``natural'' order. For those agent
classes where agents get dynamically created and
dropped (i.e.\fsp all classes other than \cat), the
ordering of the corresponding agent list will thus
suffer some degree of automatic permutation or
shuffling as the simulation proceeds; however, this
will be fairly slow.
As a general rule, in this kind of simulation, it would
be preferable to randomise the order in which agents
are updated on every timestep. This would give a
greater assurance that accidental artifacts will not
arise due to a particular update order. As can be
seen, SCL v0.04 falls somewhat short of this ideal.
However, given that there is \emph{some} randomising of
the update order (albeit relatively slow and
unsystematic), and, more importantly, that the update
order is not systematically correlated with the spatial
positions of the agents, it is probably satisfactory.
It could be improved by explicitly executing a random
permutation on each agent list on each timestep. There
is no built-in support for this in the current Swarm
libraries; however, it would not be all \emph{that}
difficult to implement, and should have only a moderate
effect on performance of the model.
With the wisdom of hindsight (!), SCL could arguably be
implemented perfectly satisfactorily on the basis of
randomly sampling agents for update (similar to a monte
carlo technique), rather than having a guaranteed
update of every agent on every (``macro'') timestep.
This would eliminate any artifacts of update order. As
an added bonus, since there would no longer be a
\texttt{ForEach:} method scanning an agent list, some of
the agent management problems might also be bypassed.
\subsection{Persistent Storage}
An essential part of the requirement for SCL was to
support persistent storage of a model---i.e.\fsp being
able to save a model into a disk file, and reload it
later. This is not supported by any of the current
Swarm example applications, and there is little
support for it built into Swarm.
Swarm does provide object classes \texttt{ObjectSaver}
and \texttt{ObjectLoader} intended to take an arbitrary
object and save it or reload it. I experimented with
these, but eventually found them too cumbersome. They
operate by exploiting the Swarm probe machinery. Thus,
in order to control exactly what instance variables are
processed, it is necessary to establish appropriate
\texttt{probeMaps}. Further it is difficult to use
these methods in a way which easily respects the class
inheritance hierarchy. Finally, in some cases,
instance variables cannot be meaningfully saved to disk
in their normal form---because they are actually
pointers, and the same object will not generally be
guaranteed to be at the same location when the model is
reloaded (indeed, it would generally be highly
unlikely). In these cases it is necessary to
``re-code'' these variables in some sense. To do this
within the \texttt{ObjectSaver} and
\texttt{ObjectLoader} framework would mean developing
special ``proxy'' objects, to which
\texttt{ObjectSaver} and \texttt{ObjectLoader} could be
applied, but which would contain the ``re-coded''
instance variables. Given all these complications, I
abandoned this approach.
The strategy I adopted was to provide all relevant
objects with methods, \texttt{saveTo:} and
\texttt{loadFrom:}, which will save to or load from a
given disk file (implemented as a Swarm
\texttt{OutFile} or \texttt{InFile} object as
appropriate). The \texttt{sclModelSwarm} then provides
methods to get a file name, open the file, and invoke
the required \texttt{saveTo:} and \texttt{loadFrom:}
methods, in order, on all required objects. This
finally worked satisfactorily, but there were various
problems along the way:
\begin{itemize}
\item The \texttt{InFile} and
\texttt{OutFile} classes did not include methods for
all native types---specifically,
\texttt{unsigned} and \texttt{BOOL}. These were saved
in \texttt{int} form instead, but this is not robust
and obscures the code.\par
\item The saving and loading methods
are not symmetrical; it is necessary to write extra
white space to guarantee correct reading.\par
\item \texttt{InFile} and \texttt{OutFile} do not record the
\emph{types} of the fields being stored, which makes it
harder to detect mismatches between the saving and
loading processes.\par
\item The \texttt{WorldRaster} class has a bug, such
that, when dropped, it does not automatically cause
the corresponding widget to be dropped.\par
\item To guarantee following exactly the same
trajectory after reloading a model, it is necessary
to store (and reload) the state of the underlying
random number generator. This is not directly
possibly with the standard Swarm generator. This is
discussed in more detail in the next section.\par
\item Similarly, to guarantee following the same
trajectory on reloading, it is necessary that the
ordering of the agent lists be preserved. This is
somewhat tricky. It means, in particular, that the
SCL model must \emph{not} be saved simply by scanning
the 2D space, and saving the particles in that order;
rather it must be done by scanning the agent lists,
and saving in \emph{that} order instead.\par
\item There is still an issue of how, exactly, to
``recode'' pointer information. In SCL the only
significant case of this was the pointers from bond
agents to \lnk{} agents and \emph{vice versa}. The
solution in this case was simply to recode the
pointers to \lnk{} agents by their world co-ordinates.
The pointers could then be reconstructed on reload
(provided, of course, that the \lnk{} particles are
reloaded, without any bond information, \emph{prior}
to the bonds). The reverse pointers, from \lnk{}
particles to bonds, can also then be re-constructed
as the forward pointers are re-established.\par
\item There is a significant issue of what, if any,
state information is hidden in the Swarm scheduling
and activity machinery. There is no built-in support
for accessing this. In the case of SCL, which does
not make use of any advanced features of this
machinery, I finessed this problem by having the only
activity be to execute the \texttt{step} method on
\texttt{sclModelSwarm}. This object (and method) are
not changed by saving and/or reloading; this
variation is all hidden in other SCL objects, which
the activity machinery is not aware of. One small
side effect of this is that the ``model time'' cannot
be taken from the Swarm activity system (as it
continues to increase monotonically, regardless of
saving and loading operations). Instead, the
\texttt{worldManager} simply keeps its own
independent record of model time, which \emph{is}
saved and/or reloaded as necessary.
\end{itemize}
\subsection{Access to Random State?}
Swarm provides a random number library, including a
variety of underlying generators. The \texttt{PMMLCG1}
generator is made available (via the globally visible
\texttt{uniformRandom} object) by default. A problem
with this arrangement is that the underlying
\texttt{PMMLCG} object does not provide access to its
internal state. This makes it impossible to stop a
simulation and save it to disk in such a way that, when
restarted, it will continue along exactly the same,
deterministic, trajectory. One may query whether this
is necessary or not: in a stochastic simulation such as
SCL, any phenomena of real interest ought to be robust
and statistically repeatable anyway. Nonetheless, for
debugging and demonstration purposes, it is often
convenient to be able to reproduce a model trajectory
exactly, and I considered that an essential feature in
SCL. I worked around this deficiency simply by
creating a new object class, \texttt{SCLRandomBit},
subclassed from \texttt{PMMLCG1}, which provided
methods to save and load the exact state of the
generator to or from a disk file. Methods are also
provided to access and/or set the state, for good
measure; in SCL these are used to allow the
\texttt{worldManager} (and thus the user) to monitor or
alter the state if desired.
\subsection{Agent Graphics}
The SCL model has five distinct classes of agents
(bonds, holes, \sbst, \cat, \lnk) which must be
displayed on a 2D raster. Bonds and \lnk{} particles
have further variations in how they should be displayed
(four bond orientations, stable versus disintegrating
\lnk{} particles, \lnk{} versus \lnkplus{} particles).
In SCL v0.04 these display issues have been addressed
in extremely crude ways. Each agent (and variation) is
represented by some form of bitmap, but these bitmaps
are generated, on the fly, pixel-by-pixel, by
dedicated, hard wired, code. This is the end point of
an incremental design process, in which the display
facilities were gradually evolved and made more
complex. The final result is functional; but it is
also extremely cumbersome and fragile (i.e.\fsp it is
difficult to modify without causing unintentional
side-effects). Performance is probably also very poor.
However: in the current release of Swarm, the only
documented methods for drawing on rasters are for
coloring individual pixels, or rectangles. This is a
significant limitation. I would hope that future
releases will provide more sophisticated 2D graphics
primitives---in particular, making it easy to display
arbitrary bitmaps. In the meantime, SCL v0.04 graphic
handling is \emph{not} a good example to emulate.
\section{Conclusion}
The artificial chemistry implemented by SCL v0.04, and
described here in detail, was qualitatively inspired by
the system described in \cite{Varela:Autopoiesis}.
However, that original \emph{algorithm} was
deliberately \emph{not} used in the development of SCL.
This arose partly due to the difficulties of
interpretation previously documented in
\cite{McMullin:OriginalAutop}; partly because I wanted
to significantly expand the parameter space of the
original model; and partly because, given the
object-oriented nature of the Swarm system, it seemed
more appropriate to develop a new, properly object
oriented, implementation.
This report has \emph{not} offered any review of the
phenomenology presented in SCL, or of any particular
experiments carried out with it; that is deferred to a
subsequent report. This report has been limited rather
to documenting the implementation of SCL and, in
particular, documenting the use of Swarm for this
purpose. I hope it may be of some value to other
researchers, possibly in quite different fields,
but interested in using the Swarm system.
I have found the Swarm system a good platform for this
development. It was especially beneficial in making it
relatively easy---via the probe mechanism---to provide
good user interface functionality. The object
oriented, agent-based, architecture proved particularly
suitable for SCL, in that the structure of the code is
able to closely mimic the structure of the qualitative
description. The Swarm release in use was still only at
beta stage, and was clearly deficient in certain
respects. Notwithstanding that, the general quality and
reliability were high. The documentation is also already
quite extensive and usable, though still incomplete.
In conclusion, it seems to me there are already good
grounds for believing that when the full, stable,
release of Swarm becomes available, it will be a very
useful and beneficial tool for scientific simulation of
agent based systems.
\nocite{Maturana:AutopoiesisAndCognition}
%\newpage
\bibliography{bmcm9702} \bibliographystyle{bmcmtr}
\newpage
\section*{Acknowledgments\label{acknowledge}}
This report was written while I was a visiting
researcher (1996/97) with the \hlnk{Swarm Project
Group}{http://www.santafe.edu/projects/swarm} at the
\hlnk{Santa Fe Institute}{http://www.santafe.edu}.
This visit was made possible by generous support from
\hlnk{Dublin City University}{http://www.dcu.ie} and
the Swarm Project. Swarm has been supported by Grant
No.~N00014-94-1-G014 from the Department of the Navy,
Naval Research Laboratory, acting in cooperation with
the Advanced Research Projects Agency. Swarm has also
benefited from earlier support from The Carol O'Donnell
Foundation and from Mr.~and Mrs.~Michael Grantham.
I am indebted to the School of Electronic Engineering in
DCU (particularly through the agency of its Head, Charles
\mbox{McCorkell}) for continuing encouragement.
All errors remain, of course, my own responsibility.
The html version of the article could not have been
created without the benefit of the {\bf LaTeX}2{\tt
HTML}\footnote{\texttt{\varurl
{http://cbl.leeds.ac.uk/nikos/tex2html/doc/
latex2html/latex2html.html}
{http://cbl.leeds.ac.uk/nikos/tex2html/doc/latex2html/latex2html.html}}}
translator developed by Nikos Drakos\footnote{\hoturl
{http://cbl.leeds.ac.uk/nikos/personal.html}} and the
Harvard\footnote{\hoturl
{http://www.arch.su.edu.au/\~{}peterw/latex/harvard/}}
extensions for author/date citations developed by Peter
Williams\footnote{\hoturl
{http://www.arch.su.edu.au/\~{}peterw/index.html}}
and Thorsten Schnier.
\section*{Revision History\label{revision}}
%\begin{quote}
%\emph{This is a draft version, circulated privately in
% November 1996. Please do not reference or
% distribute.}
%\end{quote}
First published in January 1997, simultaneously as
Working Paper No.~\sfinumber{} of the \hlnk{Santa Fe
Institute}{http://www.santafe.edu} and as Technical
Report No.~\dcunumber{} of the \hlnk{School of
Electronic Engineering}{http://www.eeng.dcu.ie} of
\hlnk{Dublin City University}{http://www.dcu.ie}.
\newpage
\section*{Author Contact Information\label{authors}}
\label{mcmullin}
\hlnk{\textbf{Barry
McMullin}}{http://www.eeng.dcu.ie/\~{}mcmullin}
\medskip
\noindent\begin{tabular}{ll}
\multicolumn{2}{l}{\hlnk{Santa Fe Institute}
{http://www.santafe.edu}}\\
\multicolumn{2}{l}{\textit{(Visiting Researcher 1996/97)}}\\
\multicolumn{2}{l}{1399 Hyde Park Road}\\
\multicolumn{2}{l}{Santa Fe, NM 87501}\\
\multicolumn{2}{l}{USA}\\[3mm]
\textbf{Voice:} & \texttt{+1-505-984-8800}\\
\textbf{Fax:} & \texttt{+1-505-982-0565}\\
\textbf{E-mail:} & \mailto{McMullin@santafe.edu}
\end{tabular}
\bigskip
\noindent\begin{tabular}{ll}
\multicolumn{2}{l}{\hlnk{School Of Electronic Engineering}
{http://www.eeng.dcu.ie}}\\
\multicolumn{2}{l}{\hlnk{Dublin City University,}
{http://www.dcu.ie}}\\
\multicolumn{2}{l}{Dublin 9,}\\
\multicolumn{2}{l}{IRELAND}\\[3mm]
\textbf{Voice:} & \texttt{+353-1-704-5432}\\
\textbf{Fax:} & \texttt{+353-1-704-5508}\\
\textbf{E-mail:} & \mailto{McMullin@eeng.dcu.ie}\\
\textbf{Web:} & \hoturl{http://www.eeng.dcu.ie/\~{}mcmullin}
\end{tabular}
\section*{Copyright\label{copyright}}
This report is copyright \copyright \thecopyrightdate{} by
\htmlref{Barry McMullin}{mcmullin}.
Permission is hereby granted to private individuals to access,
copy and distribute this work, for purposes of private study
only, provided that the distribution is complete and unmodified,
is accompanied by this copyright notice, and that no charges are
levied; multiple copying for class use in educational
organisations is specifically \emph{permitted}.
The work may \emph{not\/} be accessed or copied, in
whole or in part, for commercial purposes, except with
the prior written permission of the author.
All other rights reserved.
\newpage
\section*{Retrieval\label{retrieval}}
The resources comprising this report are electronically
retrievable, in various formats, via the \hlnk{World Wide Web}
{http://www.w3.org/hypertext/WWW/TheProject}. The root
(directory) URL is:
\medskip
\begin{center}
\noindent
\texttt{\varurl{http://www.santafe.edu/sfi/publications/
Working-Papers/97-01-002/}
{http://www.santafe.edu/sfi/publications/Working-Papers/97-01-002/}}
\end{center}
\medskip
This is also mirrored at:
\medskip
\begin{center}
\noindent\hoturl{http://www.eeng.dcu.ie/\~{}autonomy/bmcm9702/}
\end{center}
\medskip
Either of these URL's will directly access the root node
of the HTML format report. Alternatively, an
additional filename may be appended, to
retrieve one of the other formats, or related
resources, as follows:
\begin{itemize}
\item The complete set of files making up the HTML format of the
article is available in the single compressed archive file
\hoturl{bmcm9702.html.tar.gz} (c.~67~KByte),
which can be conveniently downloaded for local and/or offline
browsing.
\item A printable version of the report is available in
postscript format as file \hoturl{bmcm9702.ps}
(c.~206~KByte, 18~pages) or in compressed form as
file \hoturl{bmcm9702.ps.gz} (c.~71~KByte).
\item The original \LaTeX{} source of the report is
available as file \hoturl{bmcm9702.tex}
(c.~66~KByte) or in compressed form as file
\hoturl{bmcm9702.tex.gz} (c.~22~KByte).
\item The \BibTeX{} source for the reference
list is available as file \hoturl{bmcm9702.bib}
(c.~4~KByte).
\item A complete set of all the files just listed
is available as a single compressed
archive file \hoturl{bmcm9702.tar.gz}
(c.~162~KByte).
\item Finally, the source code distribution of SCL is available
in the single compressed archive file
\hoturl{scl-0.04.tar.gz} (c.~32~KByte)
\end{itemize}
\end{document}
 \end{rawhtml}
This is the standard Swarm \emph{control panel} window.
It provides buttons to start the simulation
(\texttt{Go}), stop it (\texttt{Stop}), single step
(\texttt{Timestep}) and terminate the simulation
(\texttt{Quit}). The buttons for controlling execution
of the simulation will, of course, operate only while a
simulation model, or world, exists. The tools for
creating and manipulating these world models will be
described separately below. This window is \emph{not}
generated as a \texttt{probeDisplay} on some object,
but is rather generated automatically as a standard
feature of Swarm.
\begin{rawhtml}
\end{rawhtml}
This is the standard Swarm \emph{control panel} window.
It provides buttons to start the simulation
(\texttt{Go}), stop it (\texttt{Stop}), single step
(\texttt{Timestep}) and terminate the simulation
(\texttt{Quit}). The buttons for controlling execution
of the simulation will, of course, operate only while a
simulation model, or world, exists. The tools for
creating and manipulating these world models will be
described separately below. This window is \emph{not}
generated as a \texttt{probeDisplay} on some object,
but is rather generated automatically as a standard
feature of Swarm.
\begin{rawhtml}
 \end{rawhtml}
This is a (tailored) \texttt{probeDisplay} on the
\texttt{parameterManager} object. It provides
facilities to monitor and modify all the model
parameters supported by SCL, and for saving and/or
reloading these parameters to or from disk files.
To modify a parameter, simply click on the relevant
position in the window and edit the value. Ensure that
you press \texttt{Enter} or \texttt{Return} to cause
the new value to take effect (this is a standard
feature of Swarm \texttt{probeDisplay} windows). Note
that, because the \texttt{probeDisplay} is continuously
refreshed while the simulation is running, it is only
practical to alter parameters while the simulation is
stopped.
Note also that no attempt has been made to make the
file interface ``robust''; specifically, any error in
accessing a file (e.g.\fsp if it does not exist and/or
cannot be created, or is of the wrong format etc.)
will result in a fatal exception being raised and SCL
will exit.
\begin{rawhtml}
\end{rawhtml}
This is a (tailored) \texttt{probeDisplay} on the
\texttt{parameterManager} object. It provides
facilities to monitor and modify all the model
parameters supported by SCL, and for saving and/or
reloading these parameters to or from disk files.
To modify a parameter, simply click on the relevant
position in the window and edit the value. Ensure that
you press \texttt{Enter} or \texttt{Return} to cause
the new value to take effect (this is a standard
feature of Swarm \texttt{probeDisplay} windows). Note
that, because the \texttt{probeDisplay} is continuously
refreshed while the simulation is running, it is only
practical to alter parameters while the simulation is
stopped.
Note also that no attempt has been made to make the
file interface ``robust''; specifically, any error in
accessing a file (e.g.\fsp if it does not exist and/or
cannot be created, or is of the wrong format etc.)
will result in a fatal exception being raised and SCL
will exit.
\begin{rawhtml}
 \end{rawhtml}
This is a (tailored) \texttt{probeDisplay} on the
\texttt{worldManager} object. It provides ``high
level'' facilities for creating and manipulating a
model world (or, more precisely, a model world
\emph{state}), and for saving and/or reloading the
world state to or from disk files.
Note that if the world state is loaded from disk then
any already loaded world model will be automatically
(and silently) dropped first. As with the file
interface for parameters, no attempt has been made to
make the world state file interface ``robust'', and any
error in accessing a file will result in a fatal
exception being raised and SCL will exit.
\texttt{PMMLCG} refers to the pseudo random number
generator object used to implement the (pseudo!)
stochastic effects in SCL. The \texttt{WorldManager}
window allows the exact state of this generator to be
monitored and/or modified. This permits, for example,
a given world state to be reloaded and then restarted
with a different initial value for the generator state,
thus potentially exploring the robustness of stochastic
phenomena in the model.
The \texttt{worldTime} records the number of full
timesteps which have elapsed. It is meaningful only
when a world model has been created or loaded from
disk.
The \texttt{WorldManager} window offers two ways of
creating a new model world. As with loading a model
from disk, if a new model is created here then any
already loaded model will be automatically (and
silently) dropped first. The
\texttt{createDefaultWorld} method provides a
``canned'' world with certain compiled-in
characteristics. The \texttt{createEmptyWorldWithX:Y:}
method allows a world of arbitrary size to be created.
This will be initially ``empty'', i.e.\fsp populated
exclusively with holes.
Once a world has been created (or loaded from disk)
particles can be incrementally added to (or removed
from) the world using the three
\texttt{adjust...NumberBy:} methods---these accept a
positive or negative integer argument, being the number
of particles to add or remove respectively. Particles
are added or removed at random positions with these
methods. They can be added only into holes; when they
are removed, they leave holes. These methods return the
number of particles \emph{actually} added or removed.
This may generally differ from the number requested.
The methods can only add new particles into existing
holes---so if the world becomes full, no more can be
added. Conversely, only existing particles can be
removed---so once all particles of a given class have
been removed no more can be taken out.
Special methods are also provided to forcibly fill the
space completely with substrate, or completely empty it
again (i.e.\fsp ``fill'' with holes).
\begin{rawhtml}
\end{rawhtml}
This is a (tailored) \texttt{probeDisplay} on the
\texttt{worldManager} object. It provides ``high
level'' facilities for creating and manipulating a
model world (or, more precisely, a model world
\emph{state}), and for saving and/or reloading the
world state to or from disk files.
Note that if the world state is loaded from disk then
any already loaded world model will be automatically
(and silently) dropped first. As with the file
interface for parameters, no attempt has been made to
make the world state file interface ``robust'', and any
error in accessing a file will result in a fatal
exception being raised and SCL will exit.
\texttt{PMMLCG} refers to the pseudo random number
generator object used to implement the (pseudo!)
stochastic effects in SCL. The \texttt{WorldManager}
window allows the exact state of this generator to be
monitored and/or modified. This permits, for example,
a given world state to be reloaded and then restarted
with a different initial value for the generator state,
thus potentially exploring the robustness of stochastic
phenomena in the model.
The \texttt{worldTime} records the number of full
timesteps which have elapsed. It is meaningful only
when a world model has been created or loaded from
disk.
The \texttt{WorldManager} window offers two ways of
creating a new model world. As with loading a model
from disk, if a new model is created here then any
already loaded model will be automatically (and
silently) dropped first. The
\texttt{createDefaultWorld} method provides a
``canned'' world with certain compiled-in
characteristics. The \texttt{createEmptyWorldWithX:Y:}
method allows a world of arbitrary size to be created.
This will be initially ``empty'', i.e.\fsp populated
exclusively with holes.
Once a world has been created (or loaded from disk)
particles can be incrementally added to (or removed
from) the world using the three
\texttt{adjust...NumberBy:} methods---these accept a
positive or negative integer argument, being the number
of particles to add or remove respectively. Particles
are added or removed at random positions with these
methods. They can be added only into holes; when they
are removed, they leave holes. These methods return the
number of particles \emph{actually} added or removed.
This may generally differ from the number requested.
The methods can only add new particles into existing
holes---so if the world becomes full, no more can be
added. Conversely, only existing particles can be
removed---so once all particles of a given class have
been removed no more can be taken out.
Special methods are also provided to forcibly fill the
space completely with substrate, or completely empty it
again (i.e.\fsp ``fill'' with holes).
\begin{rawhtml}
 \end{rawhtml}
This is a graphic image of the state of the model.
This is implemented with the Swarm \texttt{ZoomRaster}
class, and so can be magnified or reduced (by dragging
any corner). Note however that this does \emph{not}
allow ``windowing'' onto a subarea of the world model,
or panning across it: this window will always display
the entire world. If this window is resized while the
model is stopped the image will \emph{not} be
automatically updated; you can use the
\texttt{update} button in the
\texttt{WorldManager} probe to force an update.
\newpage \sbst{} particles are shown as small,
outlined, red squares; \cat{} particles are shown as
larger, filled, yellow squares. Bonds are blue line
segments. \lnk{} particles are normally shown as blue,
outlined, squares, the same size as \cat{} particles.
If an \lnk{} particle goes into the disintegrating
state then it turns grey. If an \lnk{} particle has an
absorbed substrate (i.e.\fsp it is in the \lnkplus{}
form) then this absorbed particle is still shown as a
small, outlined, red square, but internal to the \lnk{}
particle.
Moving the pointer over this window and clicking mouse
button~1 (normally the left button) will set the
co-ordinates of the \texttt{UserCellEditor}---which is
discussed in section~\ref{celleditor} below.
Moving the pointer over this window and clicking mouse
button~3 (normally the right button) will create a new
\texttt{probeDisplay} for the particle object pointed
at. These particle objects will not be detailed
further here---consult the source code to see exactly
what instance variables and methods are associated with
any particular particle class. However, it is worth
noting that this is the \emph{only} mechanism for a
user to manipulate or modify \emph{bonds}---i.e.\fsp by
creating a \texttt{probeDisplay} on an affected \lnk{}
particle, and using the relevant methods provided
there. There is \emph{no} mechanism to create a
\texttt{probeDisplay} for a bond object.
Be aware that, due to an underlying problem with the
current release of Swarm, SCL will crash if any object,
which is being probed with a \texttt{probeDisplay}, is
dropped. Since particle objects in Swarm may quite
generally get dropped, it is recommended that you only
a create \texttt{probeDisplay} on a particle object
while the simulation is stopped; and that you drop this
\texttt{probeDisplay} (using the special, red, destroy
button---\emph{not} your window manager) before
continuing execution of the model.
\begin{rawhtml}
\end{rawhtml}
This is a graphic image of the state of the model.
This is implemented with the Swarm \texttt{ZoomRaster}
class, and so can be magnified or reduced (by dragging
any corner). Note however that this does \emph{not}
allow ``windowing'' onto a subarea of the world model,
or panning across it: this window will always display
the entire world. If this window is resized while the
model is stopped the image will \emph{not} be
automatically updated; you can use the
\texttt{update} button in the
\texttt{WorldManager} probe to force an update.
\newpage \sbst{} particles are shown as small,
outlined, red squares; \cat{} particles are shown as
larger, filled, yellow squares. Bonds are blue line
segments. \lnk{} particles are normally shown as blue,
outlined, squares, the same size as \cat{} particles.
If an \lnk{} particle goes into the disintegrating
state then it turns grey. If an \lnk{} particle has an
absorbed substrate (i.e.\fsp it is in the \lnkplus{}
form) then this absorbed particle is still shown as a
small, outlined, red square, but internal to the \lnk{}
particle.
Moving the pointer over this window and clicking mouse
button~1 (normally the left button) will set the
co-ordinates of the \texttt{UserCellEditor}---which is
discussed in section~\ref{celleditor} below.
Moving the pointer over this window and clicking mouse
button~3 (normally the right button) will create a new
\texttt{probeDisplay} for the particle object pointed
at. These particle objects will not be detailed
further here---consult the source code to see exactly
what instance variables and methods are associated with
any particular particle class. However, it is worth
noting that this is the \emph{only} mechanism for a
user to manipulate or modify \emph{bonds}---i.e.\fsp by
creating a \texttt{probeDisplay} on an affected \lnk{}
particle, and using the relevant methods provided
there. There is \emph{no} mechanism to create a
\texttt{probeDisplay} for a bond object.
Be aware that, due to an underlying problem with the
current release of Swarm, SCL will crash if any object,
which is being probed with a \texttt{probeDisplay}, is
dropped. Since particle objects in Swarm may quite
generally get dropped, it is recommended that you only
a create \texttt{probeDisplay} on a particle object
while the simulation is stopped; and that you drop this
\texttt{probeDisplay} (using the special, red, destroy
button---\emph{not} your window manager) before
continuing execution of the model.
\begin{rawhtml}
 \end{rawhtml}
This is a (tailored) \texttt{probeDisplay} on the
\texttt{userCellEditor} object. It provides ``low
level'' facilities for manipulating the model world.
In particular it allows the user to ``force'' a
particle of any class at any cell in the world. It is
used by first shifting its coordinates to the desired
cell---by clicking mouse button 1 while pointing at the
cell, as mentioned above---and then forcing the desired
particle class at that cell. If you wish to modify the
particle in any further way (especially to create bonds
between \lnk{} particles) you must create a
\texttt{probeDisplay} on the particle itself and use
that, as already discussed.
\begin{rawhtml}
\end{rawhtml}
This is a (tailored) \texttt{probeDisplay} on the
\texttt{userCellEditor} object. It provides ``low
level'' facilities for manipulating the model world.
In particular it allows the user to ``force'' a
particle of any class at any cell in the world. It is
used by first shifting its coordinates to the desired
cell---by clicking mouse button 1 while pointing at the
cell, as mentioned above---and then forcing the desired
particle class at that cell. If you wish to modify the
particle in any further way (especially to create bonds
between \lnk{} particles) you must create a
\texttt{probeDisplay} on the particle itself and use
that, as already discussed.
\begin{rawhtml}